Where do we stem from? "Stammbaum" and "Stammzelle".
What is needed to make all the splendid animal life forms we see today? What are the key steps leading to humans, and what are our roots? These questions have captivated people from all walks of life, including scientists, doctors, philosophers, and scholars. Charles Darwin, a famous English naturalist, transformed our thinking about our natural history. He put forward a coherent theory for the evolution of life: In every generation, within the population of a species, there are variations, some of which may be positive and others negative. These variations may lead to some members within a population finding it easier to survive than others and are more effective in producing offspring. This process of natural selection of individuals best adapted to a given environment leads, over time, to the evolution of new life forms and species. The German biologist, artist, and philosopher Ernst Haeckel (1834-1919) was deeply fascinated by Darwin’s thought and further developed and popularised the concept of evolution – which was met by staunch opposition in various circles of the day. Haeckel worked in the University town of Jena with the support of the open-minded Dukes of Weimar, who were patrons of arts and science and attracted Friedrich Schiller and Johann Wolfgang Goethe (Fig 1). Haeckel illustrated the history of life using family trees or phylogenetic trees in German "Stammbaum" (Stamm: Stem or Phylum). This concept illustrates that all life forms have a single stem and then branch off and evolve into the staggering diversity of animals that have populated our planet in the past and present. What then sits at the very root of the "Stammbaum" and is the ancestor of all animals? Related to this concept, Ernst Haeckel proposed the "biogenetic law", which states that the development of an organism reflects a rapid and condensed version of its evolutionary history ("ontogeny recapitulates phylogeny"). This concept gave rise to the idea of a stem mother of all cells. [1, 2]. All animals develop from one single cell: the fertilised egg or zygote. Its 'germplasm' – a concept that goes back to August Weissmann before we understood that DNA is the carrier of hereditary information, is passed on from generation to generation [3]. As a result, one animal can be traced back to a single-celled zygote, and in keeping with the phylogenetic family history; all animals can be traced back to a single cell. This cell is the ultimate seed of the "Stammbaum" and our common ancestor. In his "Natürliche Schöpfungsgeschichte" (The History of Natural Creation, 1868), Haeckel coined the term we now use for cells with very special properties. Again, he used the German word for "stem" and called the cell that sits at the base of our ontogeny as well as its phylogenetic analog ancestral to all animal life the "Stammzelle" or the "stem cell."[3]

Fig 1: Ernst Haeckel coined the term 'stem cell' in the 19th century—a picture of Haeckel's study still preserved in Jena, Germany (Photographed by Ralf Jauch).
What is a stem cell?
The modern connotation has deviated from its original meaning of being the grand-grand-grand-grandmother of all cells in the animal kingdom. A stem cell has two key properties. First, it can make identical copies of itself and propagate indefinitely. In other words, a stem cell can produce more stem cells and is essentially immortal. Second, when a stem cell receives directives, it can turn into specialised cell types that serve a particular role in the body. We can distinguish stem cells by their potency. A fertilised egg is totipotent, meaning that it has the capacity to develop into a complete organism and can make a viable embryo. As cells divide, they gradually form more specialised cells. When the developing embryo implants in the womb, it contains a spherical cyst-like structure with three main cell types. In its interior, it includes a mass of pluripotent stem cells, which can form all body cells but fall short of forming cells surrounding the embryo, such as the placenta. Adults still have stem cells, but their potency is restricted to special tissues and organs. For example, neural stem cells can only form nervous system cells
The role of stem cells in animal evolution
Plants, fungi, and animals are all composed of eukaryotic cells and independently developed multicellularity. The transition from single-celled eukaryotes (protists) to complex multicellular animals has been a revolutionary move in our own evolutionary past. Our idea of how this might have occurred was influenced by studies on animals with a limited number of cell types, such as sponges, and comparisons with unicellular relatives of animals still with us today, such as choanoflagellates. Cells of unicellular organisms can temporarily change their identity and occasionally form aggregates of cells resembling multi-celled entities. Animals consist of specialised cell types that form a unit that works together in an orchestrated manner. Humans, for example, have around 200 highly specialised cells and an intricate system of organs. At their core are stem cells that control the formation of new cells. Some animals, such as placozoans or sponges, have a limited repertoire of cells and lack a nervous system or muscles. What did the unicellular stem mother of all animals look like, and which evolutionary innovations were required for it to transit to become our multicellular ancestor? It is nearly impossible for us to retrieve fossil records of these fragile entities. However, by studying which molecular toolkit nature uses today across diverse branches of the three of life, we can better understand and make inferences about which molecules were used by the cells we all stem from.
The molecules of life and their conservation
The molecular machinery that makes our cells work shows a remarkable level of conservation, which is a remarkable testament that life has a common root. The genetic code is universal, and the way genetic information is translated and copied is very similar across the domains of life. Finding the evolutionary innovations that enabled the formation of new life forms holds the key to understanding our natural history. We cannot travel back in time, and we do not have fossil records and preserved traces of genetic materials from life forms that existed 100's of millions or billions of years ago. Yet, we can compare genes of extant species and reconstruct which molecular tools our common ancestors might have used. If specific genes are unique to a particular branch of life, they might have been a unique invention along the evolutionary path leading to this branch and could have been necessary for the evolution of its ancestral species.
What do we need to turn a protist into an animal?
The game-changing event of the appearance of the first complex eukaryotic cell is thought to have happened around 2 billion years ago. After this, it took a further 1-1.5 billion years of evolutionary tinkering until the first animals emerged. Apparently, the molecular innovations required to form more complex multicellular life forms that eat plants, microorganisms or other animals were far from trivial. We now know the genome of many animals and their immediate unicellular relatives. Filastereans and choanoflagellates are unicellular water-dwelling protists most closely related to animals. Studying their life cycle and the genes they use can help us understand which leaps were needed to make an animal. Genes required to hold cells together, factors that help cells communicate and receive signals, could be prime candidates for the molecular tools exclusively used by animals and absent in their unicellular relatives. Molecular switches that turn genes on and off to define the identity of cells and program cell fate conversation during the development of animals could also be animal-exclusive evolutionary innovations. However, the deeper we investigated the gene repertoire of our unicellular relatives, the more the list of genuinely animal-specific genes and pathways shrunk. From a genetic point of view, there was not a burst of newborn genes at the dawn of animal evolution. Instead, only a small set of new genes emerged, and the overall changes to the blueprint of life that triggered the evolutionary leap to the first animal were relatively subtle.
The genes that define pluripotent stem cells
In the 1980s and 1990s, scientists learned to culture pluripotent stem cells found in mouse and human embryos in a dish and thereby had the tools to study which genes and pathways regulate the maintenance of this cell state [4 , 5]. They also found out what triggers the departure from pluripotency to all the specialised cells, such as the cells forming the skin, heart, and brain. At the core of a gene network are molecular switches that bind to specific regions in our genomic DNA and control whether genes are turned on or off. SOX2 and OCT4 are two exceptionally critical versions of such switches. SOX2 is part of a gene family whose founding member is on the Y chromosome and specifies gender. At atomic resolution, the SOX2 protein acts as a molecular wedge to bind and loosen up DNA to turn genes on [6]. OCT4 is a versatile modular protein from the so-called POU family that can shuffle around and change the way it binds to DNA and can thereby accommodate a more extensive set of DNA sequences [7]. If SOX2 and OCT4 are perturbed by mutations or by changing their abundance, pluripotent stem cells cannot function properly[8-10]. In 2006, the Japanese scientist Shinya Yamanaka made the groundbreaking discovery that we do not need to destroy embryos to obtain pluripotent stem cells. Instead, we can introduce and forcibly express four genes in differentiated cells and roll them back to an embryonic state of pluripotent stem cells [11]. Unsurprisingly, SOX2 and OCT4 were part of this magic gene cocktail that triggered a program to revert development and form what we now call induced pluripotent stem cells (iPS cells) (Fig 2). Yamanaka's findings led to a scientific earthquake, and many scientists started using this technology. We can now take cells from the skin, blood, and urine to make iPS cells from mice, humans, and many other animals. This breakthrough technology which earned Dr Yamanaka the Nobel Prize in 2012. It led to an explosion of studies on the most fundamental concepts in biology and enabled opportunities to study human disease in a completely new way, leading to a flurry of clinical trials for some of the most debilitating diseases.
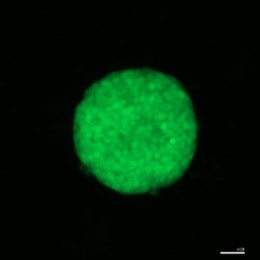
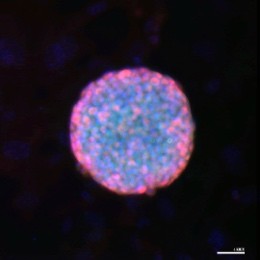
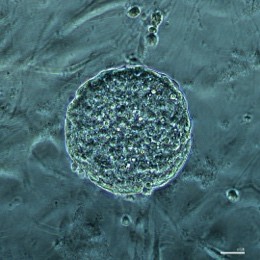
Fig 2: Microscopy images of mouse induced pluripotent stem cell (iPSC) colonies, stained with pluripotency markers, Oct4 (Green) and Sox2 (Red), captured by Dr Ya Gao.
The genes in our stem cells are older than we thought.
Our laboratory was captivated by the opportunities offered by the iPS technology. We were particularly fascinated by the molecules that drive the change of cellular identity. We sought to understand how the Yamanaka factors can direct the dramatic changes required to transform a specialised cell into an unspecialised iPS cell that proliferates and can again form all other cells of the body and even create whole mice. In our quest to discover what makes SOX2 and OCT4 so unique, we set out to delineate their evolutionary origins in collaboration with Dr Mathias Girbig and Dr Georg Hochberg of Max Planck Institute of Terrestrial Microbiology, Marburg, Germany, and Dr Alex de Mendoza of the Queen Mary University of London. The scientific literature suggested that the SOX and POU (with its prominent member of OCT4) families belong to families of genes unique to animals. Was the invention of SOX and POU genes a key step to the evolutionary transition from a unicellular protist to a multicellular animal? To investigate this question, we approached Dr Alex de Mendoza, who published influential studies on animal evolution and the gene repertoire of animals. We wanted to know what SOX and POU genes in early animals were like and how they became key players in stem cells. Dr de Mendoza revisited the question about the uniqueness of SOX and POU in animals. New protist species were discovered recently, and their gene sequences have become available. To our surprise and contrary to conventional wisdom, some of our unicellular relatives seemed to contain Sox and POU genes that matched up nicely with our SOX2 and OCT4. They previously escaped detection as they are not found in all choanoflagellate or filasterean species. Previously, only two choanoflagellate species served as scientific role models and were subject to deep inquiries. Serendipitously, these two species lack SOX and POU. Yet, less well-known members of choanoflagellates and filastereans seemed to encode genuine SOX and POU. We became very excited about this discovery and studied them in our laboratories at School of Biomedical Science of The University of Hong Kong. We wanted to compare their functions with the celebrity couple of pluripotency, SOX2, and OCT4.
700 million-year-old Ur-Sox can make mouse stem cells
We first synthesised Sox and POU genes and used techniques to produce their protein products to examine whether and how to they bind DNA. Dr Daisylyn Tan (Daisy), a talented and energetic biochemist then in the final laps of her PhD at HKUMed, took up this challenge. Daisy found that protist SOX can strongly bind the same DNA sequences as our SOX2. This led to the next question: Does this biochemical property alone allow the protist SOX to replicate the function of SOX2 to make iPS cells? This question was addressed by Dr Ya Gao, who has become an expert in stem cell biology and the intricate iPS technology during her PhD in the PACE (Protein and Cell Engineering) Laboratory (@pacelab). The ability to make iPSCs is restricted to SOX2 and its closest genetic cousins, but most other factors of the 20-strong SOX family cannot do this. Yet, amazingly, choanoflagellate Sox can induce mouse iPSCs despite 100's of million years of independent evolution. The acid test to verify the pluripotency of iPSCs is to mix them up with mouse embryos and implant them into a foster mother. To our amazement, iPS cells made with choanoflagellate SOX could contribute tissue patches to viable chimeric mice (Fig 3). After the publication of our work in Nature Communications, this patchy mouse and her iPS cell-free mate became rather famous and were featured in many media outlets, including the Washington Post and Forbes Magazine. Choanoflagellate SOX can make iPS cells as our SOX2 can do. Choanoflagellates are amongst us today, and they can look back at an evolutionary history as long as ours. Therefore, they should not be regarded as more primitive or more ancient. Ideally, we would like to travel back in time and study the molecules that existed in our past. This is precisely what Dr Georg Hochberg and Dr Mathias Girbig, evolutionary biochemists at the Max Planck Institute for Terrestrial Microbiology in Marburg, Germany, are able to do. They used the family tree made with animal and protist SOX gene sequences and used statistical models to deduce which kind of SOX genes existed in our distant ancestors. This way, they dug up ancestral Ur-SOX, which is the predecessor of SOX genes of all animals. Dr Gao showed that even this ancestral Ur-SOX can make mouse iPS cells. A hypothetical cell that was our unicellular grand-grand-grand grandmother seems to have contained a SOX factor fully equipped to unleash a genetic program required to make our modern mammalian stem cells. This ancient Ur-SOX might be older than stem cells themselves but, most conveniently, was available by the time the forces of evolution assembled the gene network that makes stem cells do their work.


Fig 3: The chimeric mouse on the left, generated from iPSCs derived from the 700-million-year-old SOX gene of choanoflagellates, exhibits black fur patches and black eyes. In contrast, the wildtype mouse on the right displays white fur and red eyes. Photos from Dr Ya Gao and Alvin Kin Shing Lee (CCMR). From our publication [12]
Were changes to POU genes a key event on the way to animals?
The evolutionary history of POU genes is more complicated and dynamic. Dr Alex de Mendoza's gene hunt in lesser-known relatives of animals also revealed that these genes must have existed before the first animals roamed our planet. However, Daisy's careful biochemical analyses showed that choanoflagellate POU binds DNA very differently than the stem cell factor OCT4. Unsurprisingly, choanoflagellate POU cannot substitute for OCT4 to generate mouse iPS cells. On the evolutionary path to animals, POU genes have likely undergone steps of evolutionary adaptations. We do not know when and how they happened, but we surmise that the first animal had a POU factor that bound DNA and regulated genes differently from choanoflagellate POU. Whether SOX and POU factors have already formed an intimate partnership in the stem mother of all animals, as SOX2 and OCT4 do in our pluripotent stem cells, is a fascinating albeit highly speculative idea. If so, evolution might have taken advantage of two pre-existing factors to select master regulators of stem cells. Ur-SOX was ready long before the first animal showed up to be incorporated into this network, while Ur-POU required more tinkering on the way to animals to take up this role.
Can we learn from evolution to engineer stem cells to cure diseases and slow ageing?
Learning about the history of life and the evolution of the molecular building blocks that shape the world around us is captivating in itself. Beyond academic fascination, the lessons from nature can also provide us with the tools to tackle some of the burdens of humankind. Scientists have for some time applied mechanisms of evolution to optimise and functionally change proteins in the laboratory. We have tools to enforce and accelerate the process of the targeted or random generation of mutations. We can design rigorous experiments to select the 'fittest' protein variants that serve a desired purpose. We can do this in many successive repeats, thereby gradually evolving proteins in the laboratory. Laboratory engineered designer proteins created in this way now have countless applications in biotechnology, agriculture, are used as drugs, and therapeutic supplements. We can also use designer proteins to turn body cells into stem cells more quickly and more efficiently. Recently, the artificial intelligence start-up Retro Biosciences [13] supported by the OpenAI founder Sam Altmann with 180M $USD, has joined the party and reported artificial versions of SOX2 and KLF4. Their artificial intelligence algorithms were trained by comparing many protein sequences and learning from their natural evolution. The hope to use their AI modified proteins to reverse age-linked diseases. Our laboratory has been using laboratory evolution for many years to boost the production of various stem cells. We believe that re-engineered proteins can help us to study ageing, model diseases, enable species conservation, and produce stem cells of higher quality. We can produce stem cells that can be turned into a broader array of tissues and even mini-organs. We hope to learn to make old stem cells to have authentic cells to capture what goes wrong in age-linked diseases. We may even be able to use cell reprogramming to change the identity of cells in our body to repair damaged organs. Some scientists have bold hopes that we can slow down or even halt the burden of ageing. The Yamanaka factors and related molecules play key roles in the efforts to manipulate cells in the laboratory and inside the body of animals. Learning from evolution to travel back in time to resurrect ancient proteins and using this as a guide to make entirely new proteins from scratch will help us optimise the factors that make the cells we need to study ageing and eventually cure diseases.
Fig 4: Finding the roots of animal evolution with the ancestral Ur-Sox (Drawn by Martha Shi Wing Yeung)
Original Publication:
Gao, Y., Tan, D. S., Girbig, M., Hu, H., Zhou, X., Xie, Q., Yeung, S. W., Lee, K. S., Ho, S. Y., Cojocaru, V., Yan, J., Hochberg, G. K. A., de Mendoza, A., & Jauch, R. (2024). The emergence of Sox and POU transcription factors predates the origins of animal stem cells. Nat Commun, 15(1), 9868. https://doi.org/10.1038/s41467-024-54152-x
References:
- Maehle, A.H., Ambiguous cells: the emergence of the stem cell concept in the nineteenth and twentieth centuries. Notes Rec R Soc Lond, 2011. 65(4): p. 359-78.
- Levit, G.S., et al., The biogenetic law and the Gastraea theory: From Ernst Haeckel's discoveries to contemporary views. J Exp Zool B Mol Dev Evol, 2022. 338(1-2): p. 13-27.
- Droscher, A., Images of cell trees, cell lines, and cell fates: the legacy of Ernst Haeckel and August Weismann in stem cell research. Hist Philos Life Sci, 2014. 36(2): p. 157-86.
- Evans, M.J. and M.H. Kaufman, Establishment in culture of pluripotential cells from mouse embryos. Nature, 1981. 292(5819): p. 154-6.
- Thomson, J.A., et al., Embryonic stem cell lines derived from human blastocysts. Science, 1998. 282(5391): p. 1145-7.
- Tan, D.S., et al., SOX17 in cellular reprogramming and cancer. Semin Cancer Biol, 2020. 67(Pt 1): p. 65-73.
- Malik, V., D. Zimmer, and R. Jauch, Diversity among POU transcription factors in chromatin recognition and cell fate reprogramming. Cell Mol Life Sci, 2018. 75(9): p. 1587-1612.
- Masui, S., et al., Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat Cell Biol, 2007. 9(6): p. 625-35.
- Tapia, N., et al., Dissecting the role of distinct OCT4-SOX2 heterodimer configurations in pluripotency. Sci Rep, 2015. 5: p. 13533.
- Malik, V., et al., Pluripotency reprogramming by competent and incompetent POU factors uncovers temporal dependency for Oct4 and Sox2. Nat Commun, 2019. 10(1): p. 3477.
- Takahashi, K. and S. Yamanaka, Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell, 2006. 126(4): p. 663-76.
- Gao, Y., et al., The emergence of Sox and POU transcription factors predates the origins of animal stem cells. Nat Commun, 2024. 15(1): p. 9868.
- RetroBiosciences. 2025; Available from: https://www.retro.bio/.
Author:
Prof Ralf Jauch, Professor, School of Biomedical Sciences, The University of Hong Kong ; Principal Investigator, Centre for Translational Stem Cell Biology
Acknowledgement
We thank Dr Mathias Girbig, Dr Alex de Mendoza, Dr Daisylyn Tan, Dr Ya Gao and Hannah Lam for critical feedback on this feature article.
March 2025
