Epidemiological Study on Coronavirus 2019 at The Hong Kong Polytechnic University
Coronavirus 2019-20 (COVID-19) pandemic has already caused tremendous impact on global public health, world economy and socioeconomic landscape. A research team from The Hong Kong Polytechnic University, formed at the onset of the outbreak, has been involved in the forefront of the epidemiological study of COVID-19. The team includes Dr He Daihai and Dr Lou Yijun from the Department of Applied Mathematics and Dr Yang Lin from the School of Nursing at The Hong Kong Polytechnic University. Since the start of the epidemic, which later became a pandemic, the team has worked tirelessly on data analyses and modeling analyses of the COVID-19 with the aim to inform policymakers and the public with updated accurate information.
In summary, the team is one of the most active epidemiological research teams on COVID-19 and has made some insightful findings which helped inform the public and policymakers. The publication prepared by the team on the preliminary estimation of the basic reproduction number of COVID-19 in Wuhan, Hubei, China in the early phase of the outbreak1 has been cited 418 times on Google Scholar, which is an indicator of the importance of the work.
Here is a short extract of its work. For details of the various research findings, please refer to the references at the end of the article.
Key Findings on Research Conducted by The Hong Kong Polytechnic University
Accurate estimate of the key epidemiological parameters2 of COVID-19 is crucial in mitigation of the outbreak. We, the team from The Hong Kong Polytechnic University, are among the first few teams to reach the conclusion on 23 January 2020 that the transmissibility (the basic reproductive number) of the COVID-19, i.e., on average one primary case may cause 2-3 secondary cases. We relied on publicly available data. It is worth mentioning that that we are the first team to point out the impact of ascertainment rate (reporting rate, reporting effort) on the estimate of basic reproductive number1. Part of the sudden increase in daily new cases is due to improved testing capacity. In a later work, we quantified the under-reported cases in middle January in Wuhan due to low awareness and inadequate testing capacity3. These tasks are very important for understanding the scale of the epidemic.
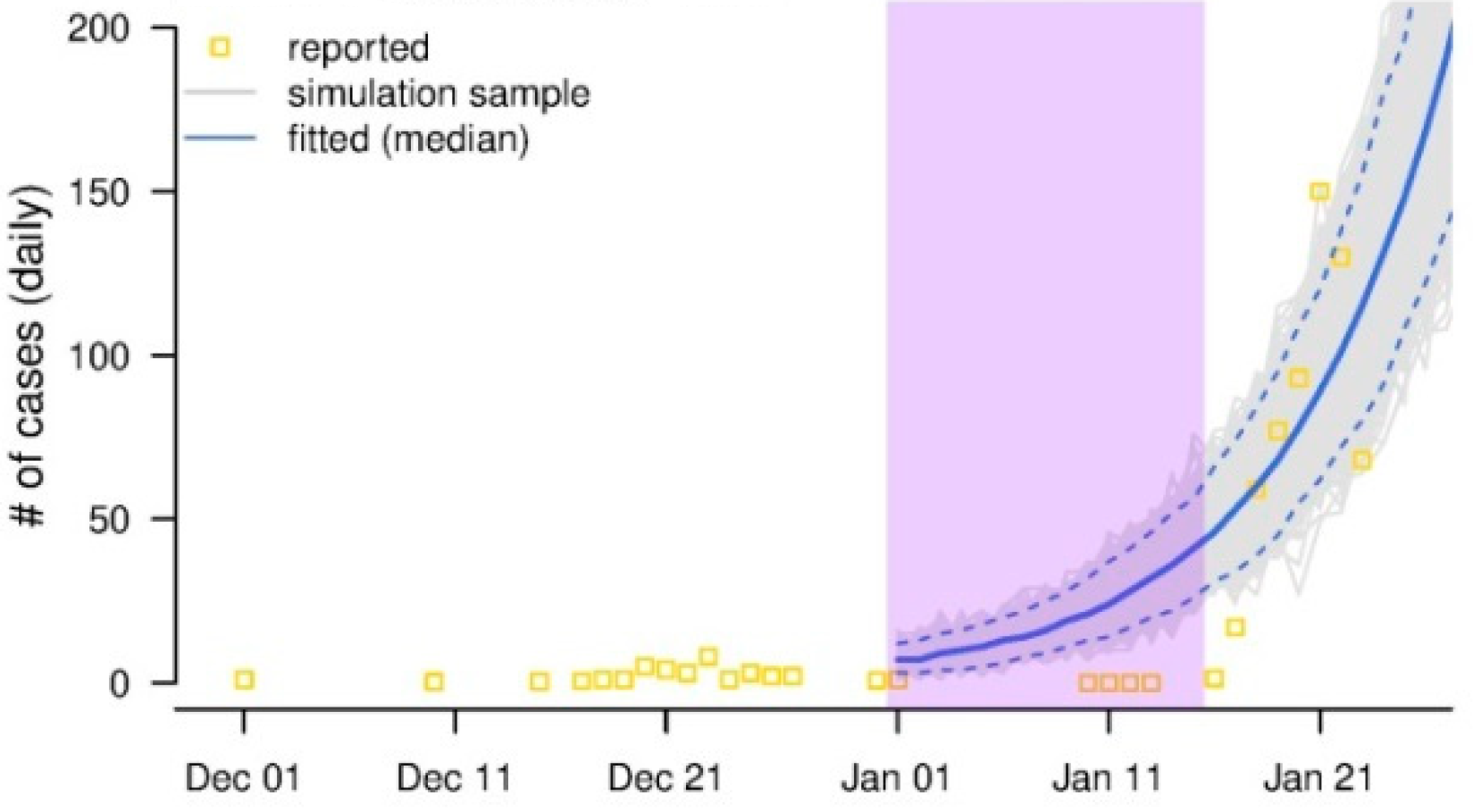
Figure. The yellow squares denote the daily reported new cases. Classic epidemiology theory tells us the daily new cases will grow exponentially in an outbreak, as illustrated by the blue curve. We noticed that in the time interval shaded by purple colour, the daily new cases are most likely being under-reported. By comparing the blue curve (theory prediction) and the actual reported cases, we estimate the under-reported cases for the purple time interval3.
At the same time, we quantified the impact of transportation4, 5, (especially via train) on the spatial spread of COVID-19 from Wuhan to other mainland cities. We obtained reported cases in cities other than Wuhan and the population flow from Wuhan to other cities, using simple linear regression model, we estimated the impact of transportation on exportation of cases from Wuhan to other cities. We also quantified the relative transmissibility of the asymptomatic COVID-19 cases6. We found that the transmissibility of asymptomatic COVID-19 cases is 1/4 of that of symptomatic cases. Thus, the asymptomatic COVID-19 cases may play a weaker role in the spreading of COVID-19.
We also calculated the serial interval7, 8 (time delay from symptom onset of primary case to symptom onset of secondary case), which is an important parameter, and together with the basic reproductive number, controls the spread speed of COVID-19. The basic reproductive number determined the expected number of secondary cases per primary case, whereas, shorter serial interval of COVID-19 implies that secondary cases emerge quickly once there is a primary case. Therefore, timely identification and isolation of patients are essential.
We are the first to quantify the variation in the transmissibility of COVID-19 cases9. We found that the variation of COVID-19 is significantly smaller than that of Severe Acute Respiratory Syndrome (SARS) which partly explained why COVID-19 is likely to stay and difficult to eradicate. During the SARS outbreak in Hong Kong in 2003, infectiousness of patients showed large variations, i.e., a few were very infectious (so called super-spreader), thus caused many secondary cases, but most were not very infectious, thus caused few secondary cases. This made SARS relatively easy to control, i.e., targeting super-spreaders. While in COVID-19, we noticed there were super-spreader events, but the infectiousness of patients was relatively even, thus it is easy for the COVID to persist in a population without strict control measure.
We developed mathematical models to assess the impact of control measures and forecast the trend of the outbreak in Wuhan, China10, 11. We compared the 1918-19 influenza pandemic with COVID 2019-20 pandemic.12 Given the similar in severity of transmissibility of 1918-19 influenza and COVID-19, we argued that the multiple-wave feature and duration of the former could be used as reference for the latter.
Jointly with Shenzhen Center for Disease Control and Prevention, we quantified the relapse rate (the proportion of discharged/recovered COVID-19 patients showing RT-PCR testing in follow-up tests) in Shenzhen13. COVID-patients are discharged if two separate RT-PCR tests are negative and no symptoms. However, we found that 10.5% discharged patients showed positive RT-PCR tests around an average of 4.7 days after discharge. This may partly due to low testing accuracy and/or complicated in-host viral dynamic. We are the first to quantify the rate (frequency of such cases will occur) and the related risk factors.
Dr Yang and collaborators also published work on maternal and neonatal outcomes of pregnant women with COVID-19 pneumonia14, and mental health, risk factors, and social media use during the COVID-19 epidemic and cordon sanitaire among the community and health15. These are very important topics.
Our work is supported by Research Grant Council of Hong Kong General Research Grant and Alibaba (China) Collaborative research grant.
References
Selected COVID-19 publications and preprints of Dr He Daihai, Dr Yang Lin and Dr Lou Yijun
- Zhao S, Lin Q, Ran J, Musa SS, Yang G, Wang W, Lou Y, Gao D, Yang L, He D, and Wang MH. (2020) Preliminary estimation of the basic reproduction number of novel coronavirus (2019-nCoV) in China, from 2019 to 2020: A data-driven analysis in the early phase of the outbreak. International Journal of Infectious Diseases. 01-050.
- Li Ying-ke, Zhao Shi, Lou Yi-jun, Gao Dao-zhou, Yang Lin, He Dai-hai (2020) Epidemiological parameters and models of coronavirus disease 2019 (COVID-19). Acta Physica Sinica, accepted.
- Zhao S, Musa SS, Lin Q, Ran J, Yang G, Wang W, Lou Y, Yang L, Gao D, He D, and Wang MH. (2020) Estimating of the unreported number of novel coronavirus (2019-nCoV) cases in China in the first half of January 2020: A data-driven modelling analysis of the early outbreak. Journal of Clinical Medicine. 9(2), 388.
- Zhao S, Zhuang Z, Cao P, Ran J, Gao D, Lou Y, Yang L, Cai Y, Wang W, He D, and Wang M. (2020) Quantifying the association between domestic travel and the exportation of novel coronavirus (2019-nCoV) cases from Wuhan, China in 2020: A correlational analysis. Journal of Travel Medicine. 2020:1-3.
- Zhao S, Zhuang Z, Ran J, Lin J, Yang G, Yang L and He D. (2020) The association between domestic train transportation and novel coronavirus outbreak in China, from 2019 to 2020: A data-driven correlational report. Travel Medicine and Infectious Diseases. 33:101586.
- He D, Zhao S, Lin Q, Zhuang Z, Cao P, Wang MH, and Yang L (2020). The relative transmissibility of asymptomatic COVID cases among close contacts. International Journal of Infectious Diseases. In press.
- Zhao S, Cao P, Gao D, Zhuang Z, Cai Y, Ran J, Chong MKC, Wang K, Lou Y, Wang W, Yang L, He D, and Wang MH (2020) Serial interval in the estimation of reproduction number of the novel coronavirus disease (COVID-19) during the early outbreak. Journal of Travel Medicine. In press.
- Zhao S, Cao P, Chong MKC, Gao D, Lou Y, Ran J, Wang K, Wang W, Yang L, He D, and Wang MH. (2020) COVID-19 and gender-specific difference: Analysis of public surveillance data in Hong Kong and Shenzhen, China, from January 10 to February 15, 2020. Infection Control & Hospital Epidemiology. In press.
- He D, Zhao S, Xu X, Zhuang Z, Cao P, Wang M H, Lou Y, Wu Y & Yang L (2020). Individual variation in infectiousness of coronavirus 2019 implies difficulty in control. Available at SSRN 3559370.
- Lin Q, Zhao S, Gao D, Lou Y, Yang S, Musa SS, Wang MH, Cai Y, Wang W, Yang L, He D. (2020) A conceptual model for the outbreak of coronavirus disease 2019 (COVID-19) in Wuhan, China with individual reaction and governmental action. International Journal of Infectious Diseases. 93:211-216.
- Zhao S, Stone L, Gao D, Musa SS, Chong MKC, He D, and Wang MH (2020) Imitation dynamics in the mitigation of the novel coronavirus disease (COVID-19) outbreak in Wuhan, China from 2019 to 2020. Annals of Translational Medicine. In press.
- [1] He D, Zhao S, Li Y, Zhuang Z, Cao P, Gao D, ... & Yang L (2020). History is the best model–a comparison of the COVID-19 with the 1918-19 pandemic influenza in United Kingdom. Available at SSRN 3582393.
- Tang X, Zhao S, He D, Yang L, Wang MH, Li Y, Mei S & Zou X (2020). Positive RT-PCR tests among discharged COVID-19 patients in Shenzhen, China. Infection Control & Hospital Epidemiology1-7.
- Li N, Han L, Peng M, Lv Y, Ouyang Y, Liu K, ... & Yang L (2020). Maternal and neonatal outcomes of pregnant women with COVID-19 pneumonia: a case-control study. Clinical Infectious Diseases.
- Ni M Y, Yang L, Leung C M, Li N, Yao X I, Wang Y, ... & Liao Q (2020). Mental health, risk factors, and social media use during the COVID-19 epidemic and cordon sanitaire among the community and health professionals in Wuhan, China. JMIR Public Health and Surveillance.
May 2020